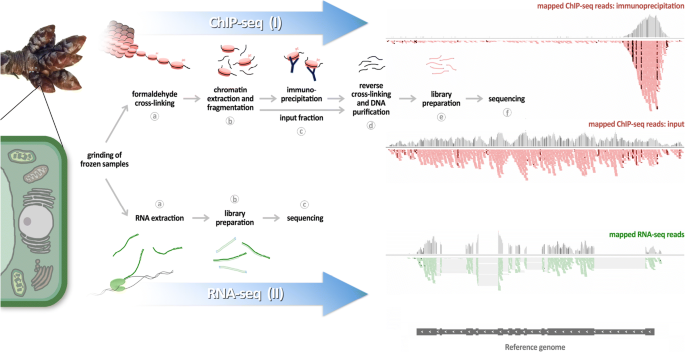
ChIP-seq protocol for sperm cells and embryos to assess environmental impacts and epigenetic inheritance - ScienceDirect
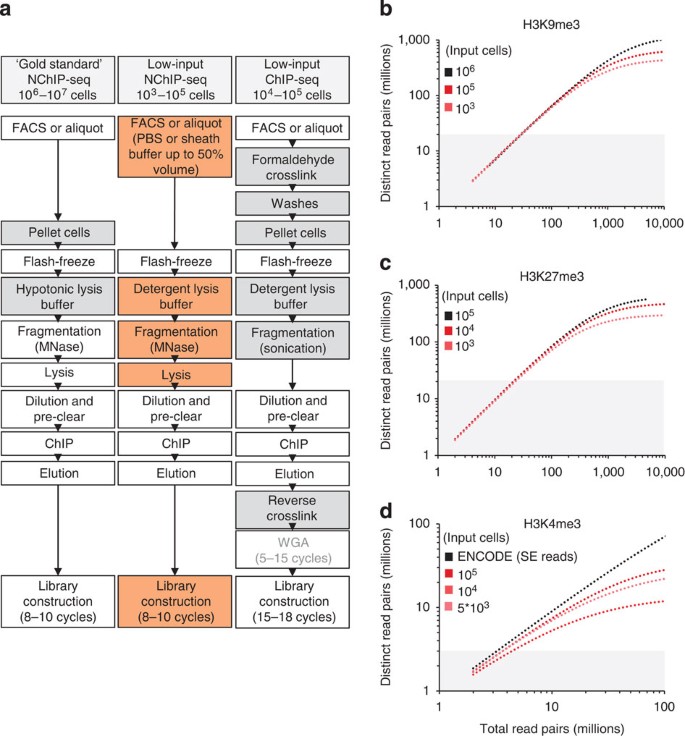
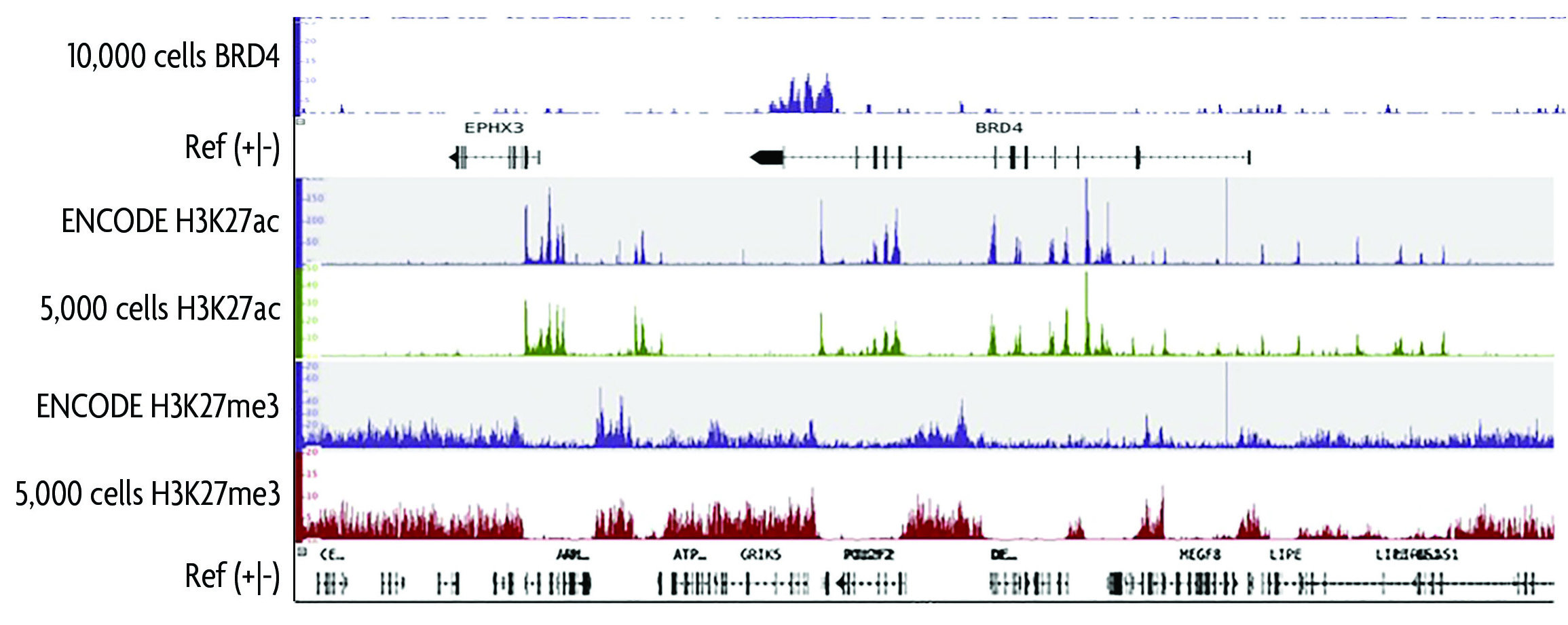
An ultra-low-input native ChIP-seq protocol for genome-wide profiling of rare cell populations | Nature Communications
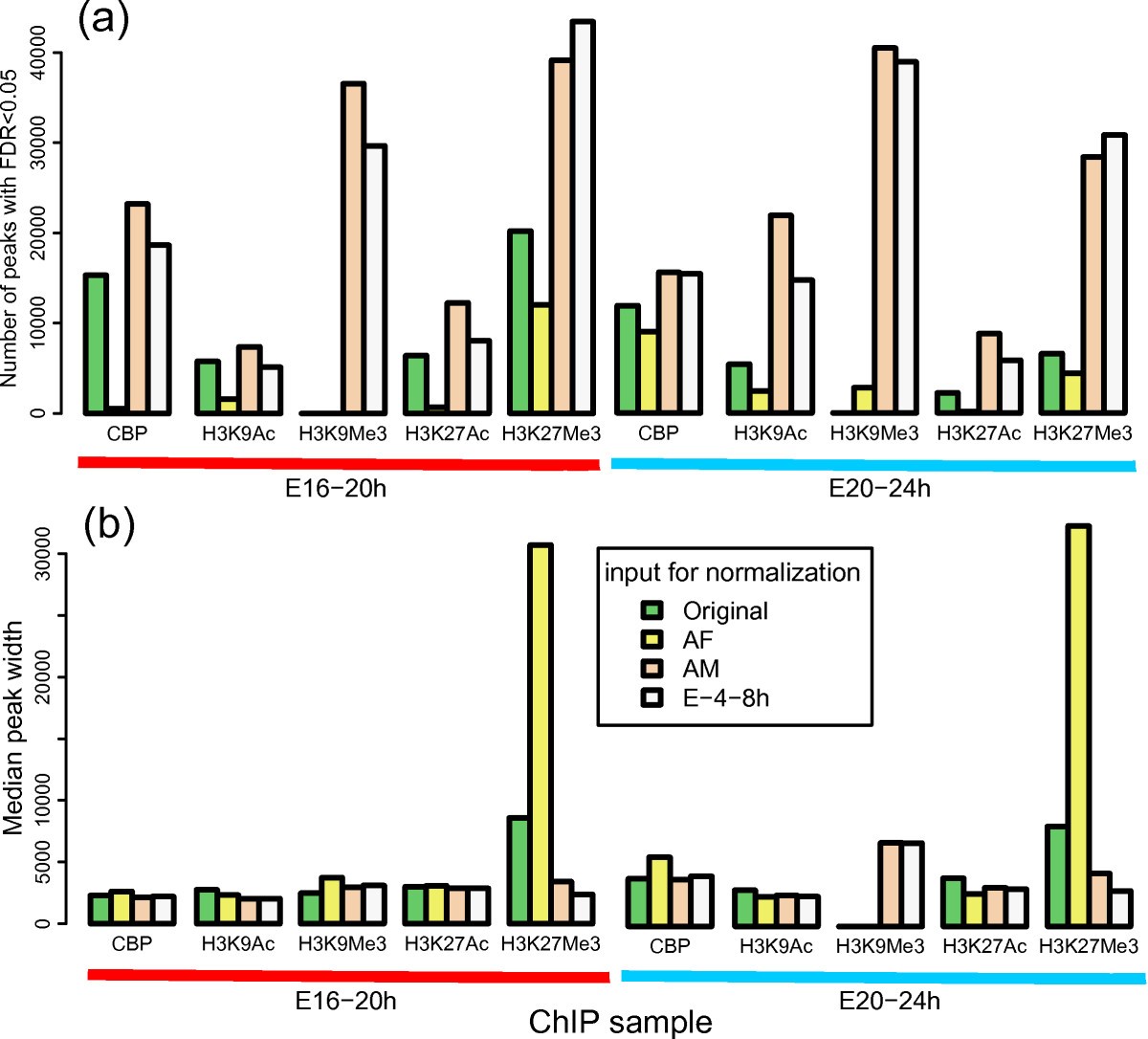
ChIP-chip versus ChIP-seq: Lessons for experimental design and data analysis | BMC Genomics | Full Text

An optimized protocol for rapid, sensitive and robust on-bead ChIP-seq from primary cells: STAR Protocols

TAF-ChIP: an ultra-low input approach for genome-wide chromatin immunoprecipitation assay | Life Science Alliance
ChIP-seq and In Vivo Transcriptome Analyses of the Aspergillus fumigatus SREBP SrbA Reveals a New Regulator of the Fungal Hypoxia Response and Virulence | PLOS Pathogens
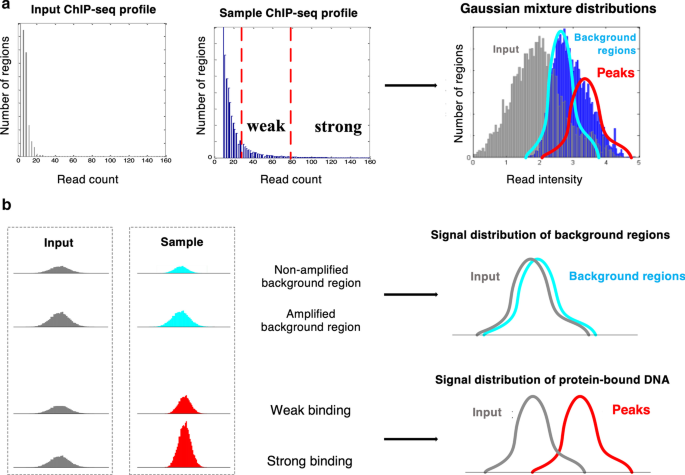
ChIP-BIT2: a software tool to detect weak binding events using a Bayesian integration approach | BMC Bioinformatics | Full Text